Research
Exploring the intersection of artificial intelligence, robotics, and human behavior through computational approaches. My research focuses on developing adaptive systems that can learn, reason, and interact effectively in complex environments.
Infra-Bayesian Reinforcement Learning Agents Outperform Classical RL For Worst-Case Robustness
Supervised Program for Alignment Research (SPAR)
An infra-Bayesian reinforcement learning agent that tracks a set of plausible world models instead of a single posterior, and selects actions by their worst-case expected value across that set.
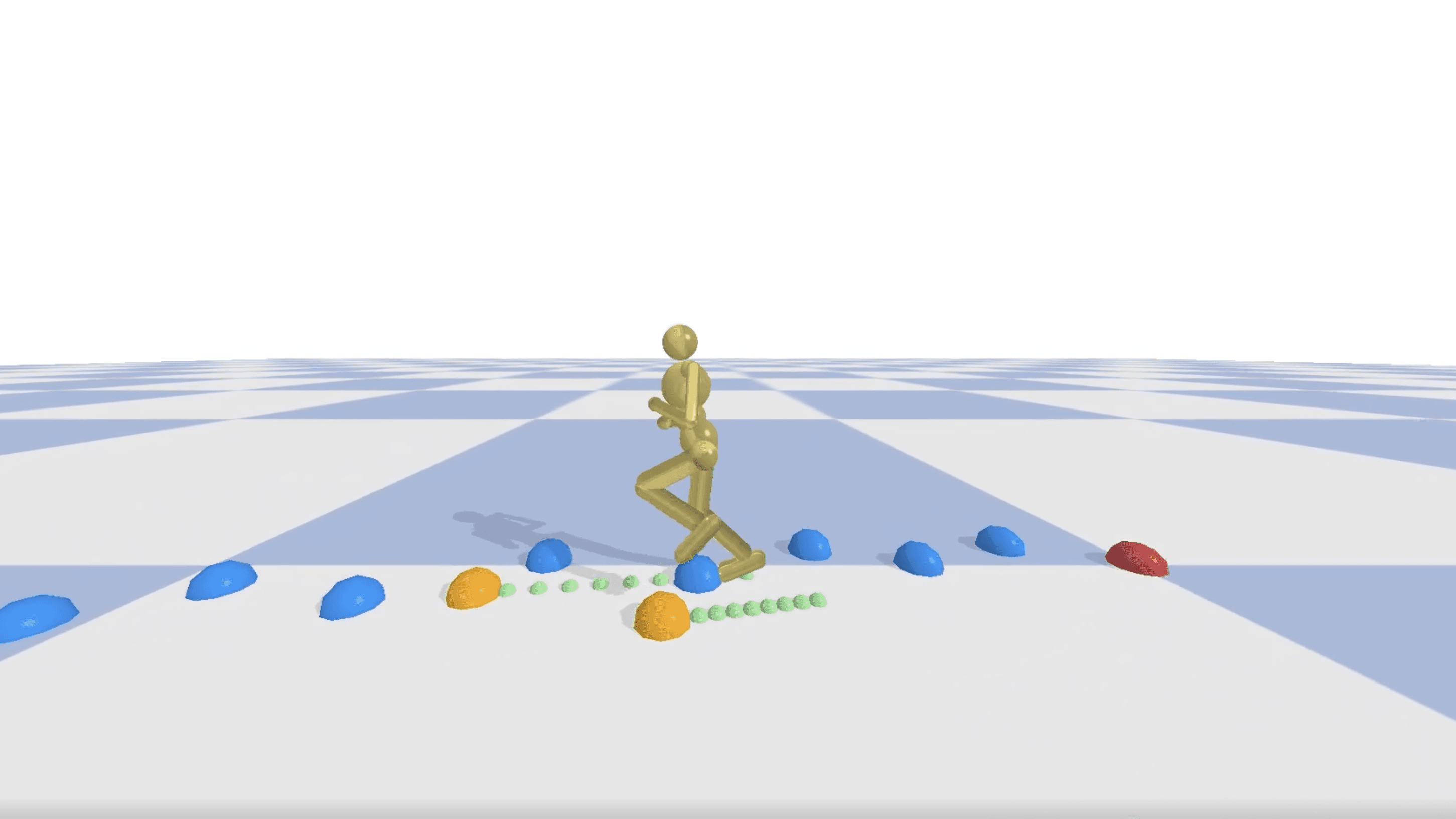

Imitation-Free Diffusion Policy Training for Humanoid Footstep Planning
MOCCA Lab, UBC Computer Science · Dr. Michiel Van de Panne
Developing robust control policies for humanoid robots to navigate challenging terrains using deep reinforcement learning techniques.
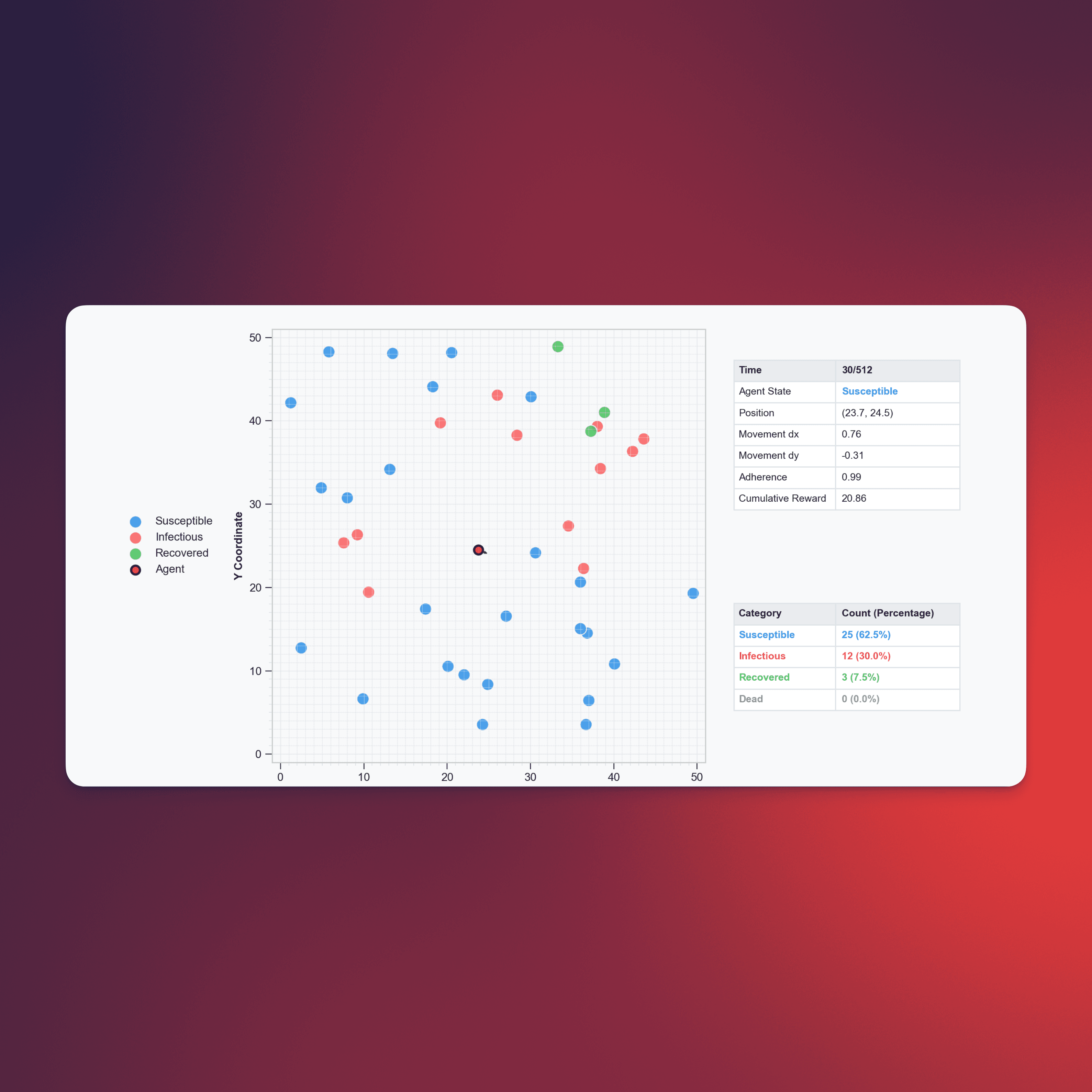
ContagionRL: A Flexible Platform for Learning in Different Spatial Epidemic Environments
UBC Mathematics & Computer Science · Dr. Daniel Coombs
ContagionRL simulates human behavioral responses during epidemics using reinforcement learning, combining a spatial SIRS disease model with single-agent RL.
Entrepreneurship Education Research
UBC Computer Science and Sauder School of Business · Dr. Angele Beausoleil
An NLP-based system that automatically analyzes and maps entrepreneurship education programs and course syllabi against defined competency frameworks using zero-shot classification.
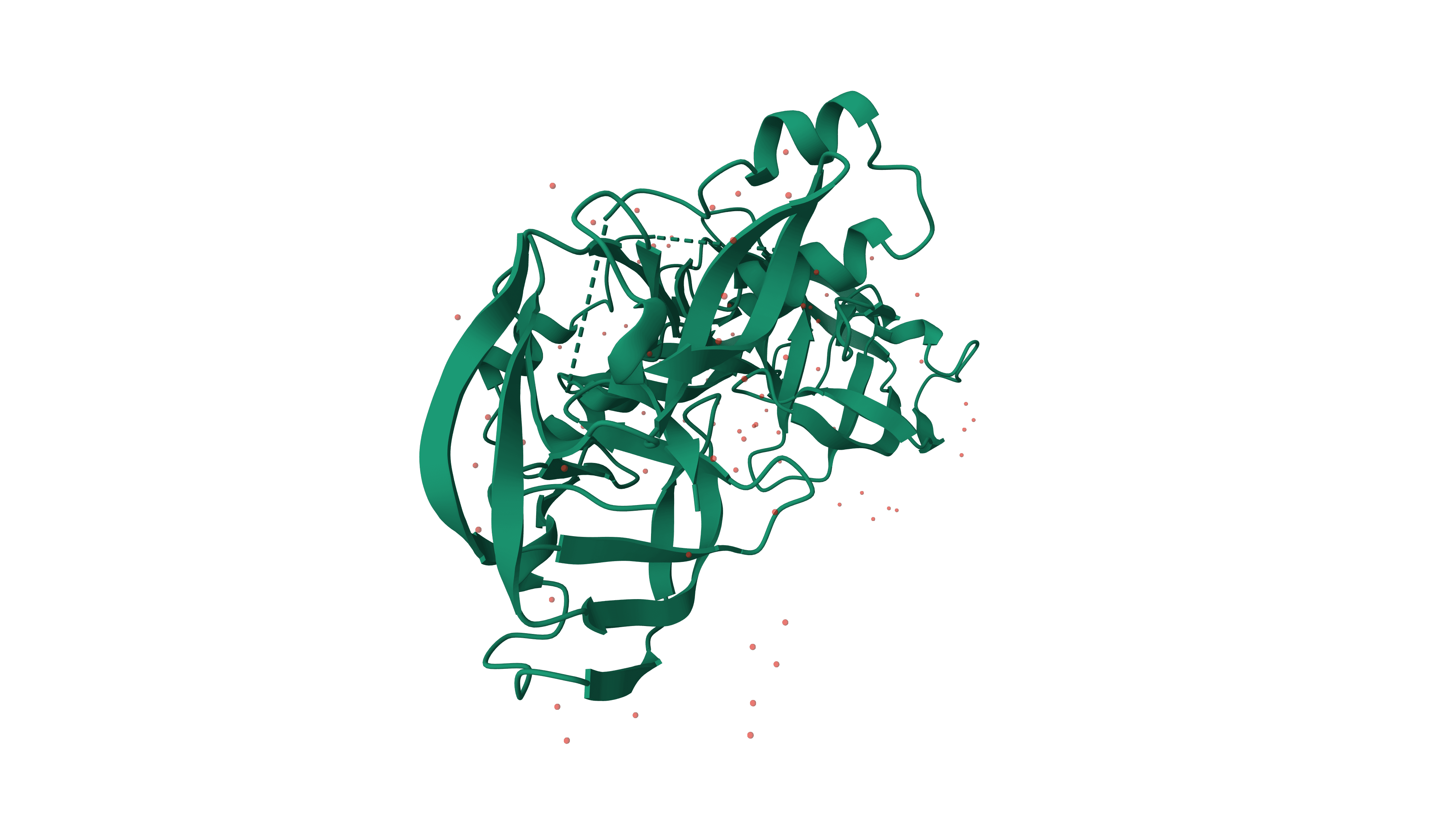
PILOT: Platformed Inteins: A linked orthogonal toolkit
UBC Life Sciences Institute (LSI) · Dr. Steven Halem
A modular intein-mediated cell-free protein synthesis platform with a self-aggregating solubility tag, enabling multisubunit peptide assembly and traceless purification in a Vibrio natriegens lysate.